In a paper published today in the Journal of Synchrotron Radiation, to which I collaborated while a postdoc in IPANEMA, we developed a robust framework and software implementation for fast speciation mapping.
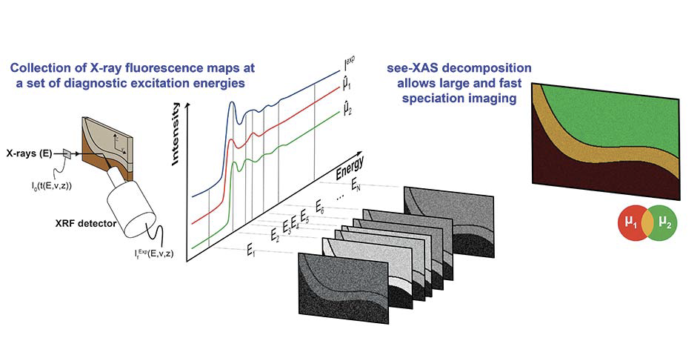
One of the greatest benefits of synchrotron radiation is the ability to perform chemical speciation analysis through X-ray absorption spectroscopies (XAS). XAS imaging of large sample areas can be performed with either full-field or raster-scanning modalities. A common practice to reduce acquisition time while decreasing dose and/or increasing spatial resolution is to compare X-ray fluorescence images collected at a few diagnostic energies. Several authors have used different multivariate data processing strategies to establish speciation maps.
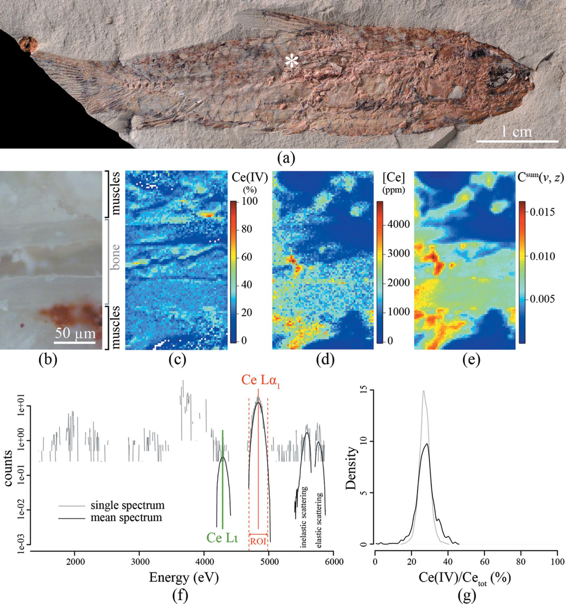
In this manuscript, the theoretical aspects and assumptions that are often made in the analysis of these datasets are presented. A robust framework (algorithm) is developed to perform speciation mapping in large bulk samples at high spatial resolution by comparison with known references. We also provide two fully operational software implementations: a user-friendly implementation within the MicroAnalysis Toolkit software, and a dedicated script developed under the R environment. The procedure is exemplified through the study of a fossil sample.
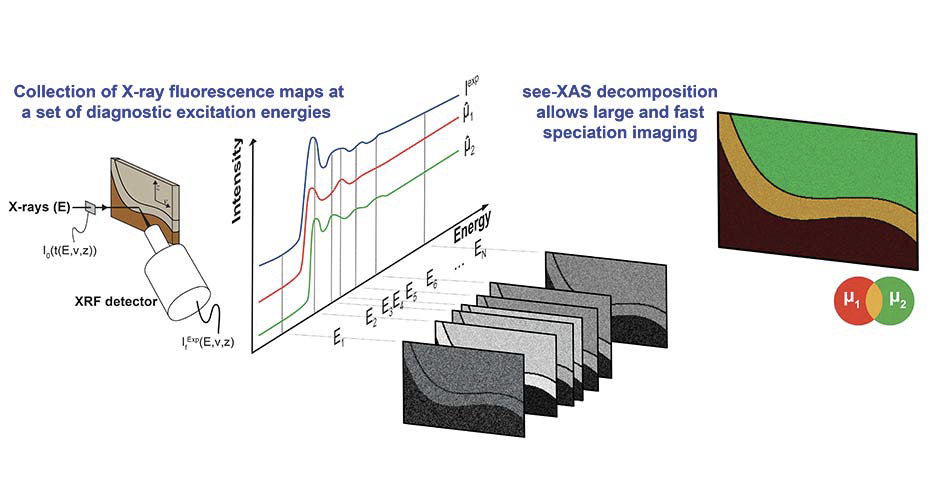
The algorithm provides accurate speciation and concentration mapping while decreasing the data collection time by typically two or three orders of magnitude compared with the collection of whole spectra at each pixel. Whereas acquisition of spectral datacubes on large areas leads to very high irradiation times and doses, which can considerably lengthen experiments and generate significant alteration of radiation-sensitive materials, this sparse excitation energy procedure brings the total irradiation dose greatly below radiation damage thresholds identified in previous studies. This approach is particularly adapted to the chemical study of heterogeneous radiation-sensitive samples encountered in environmental, material, and life sciences.
Reference: Cohen S.X., Webb S.M., Gueriau P., Curis E. & Bertrand L. 2020. Robust framework and software implementation for fast speciation mapping. Journal of Synchrotron Radiation 27: 1049–1058. Find the article here